 Back to tools
Back to tools
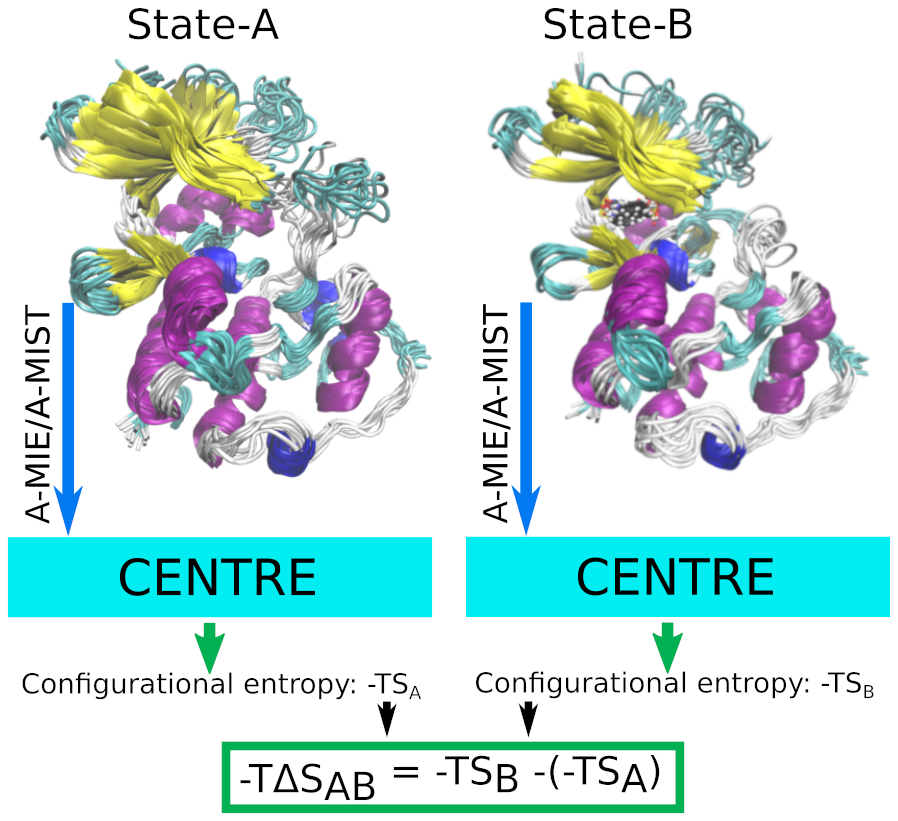
CENTRE
- Configurational ENTRopy Estimation Tool
- People: Indira Ghosh , Shailesh Kumar Panday
The CENTRE is a parallel and scalable tool for estimating Configurational entropy from MD simulation data using Information Theoretic methods:
- Mutual Information Expansion (MIE)
- Maximum Information Spanning Tree (MIST)
- Approximate Mutual Information Expansion (or Neighbor Approximated MIE i.e. A-MIE)
- Neighbor Approximated Maximum Information Spanning Tree (A-MIST)
This program accepts BOND/ANGLE/TORSION (BAT) format trajectory of the system for which the entropy is to be computed. An auxilary program cart2bat shipped with pycentre (https://github.com/shaileshp51/pycentre) python package can be used to generate a BAT format file from topology and trajectory files generated using popular Molecular Dynamics programs e.g. AMBER, NAMD, CHARMM, ACEMD, GROMACS.
See more details at https://github.com/shaileshp51/centre
Citation(s)
SK Panday and I Ghosh. "Application and Comprehensive Analysis of Neighbor Approximated Information Theoretic Configurational Entropy Methods to Protein–Ligand Binding Cases", J. Chem. Theory Comput. 2020, DOI: 10.1021/acs.jctc.0c00764
